Histone H3K36me3 (Tri-methy Lys36) antibody - ChIP grade
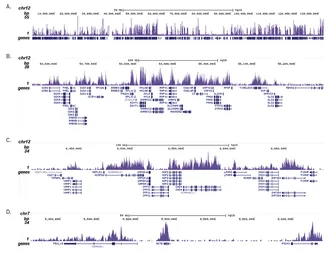
ChIP analysis of sheared chromatin from 1x10⁵ K562 cells using GTX60817 Histone H3K36me3 (Tri-methy Lys36) antibody - ChIP grade. The IP'd DNA was subsequently analysed on an Illumina Genome Analyzer. Library preparation, cluster generation and sequencing were performed according to the manufacturer's instructions. The 36 bp tags were aligned to the human genome using the ELAND algorithm. Figure 3 shows the H3K36me3 signal distribution along the complete sequence and a zoomin of human chromosome 12 (figure 2A and B) and in 2 genomic regions containing the GAPDH and ACTB positive control genes (figure 3C and D).
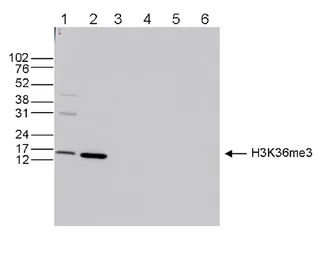
Antibody amount : 0.5μg
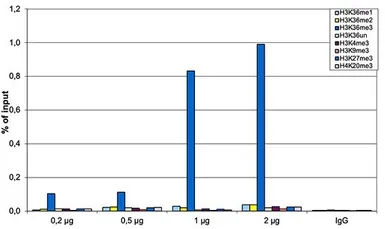
ChIP analysis of sheared chromatin from 10⁶ HeLa cells using GTX60817 Histone H3K36me3 (Tri-methy Lys36) antibody - ChIP grade. The chromatin was spiked with a panel of in vitro assembled nucleosomes, each containing a specific lysine methylation (SNAP-ChIP K-MetStat Panel, Epicypher). A titration consisting of 0.2, 0.5, 1 and 2 μg of antibody per ChIP experiment was analyzed. IgG (1 μg/IP) was used as a negative IP control. Quantitative PCR was performed with primers specific for the nucleosomes carrying the H3K36me1, H3K36me2, H3K36me3, H3K4me3, H3K9me3, H3K27me3 and H4K20me3 modifications and the unmodified H3K4. The graph shows the recovery, expressed as a % of input. These results demonstrate a high specificity of the H3K36me3 antibody for the modification of interest.
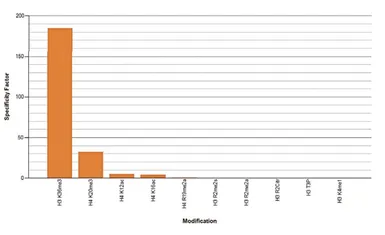
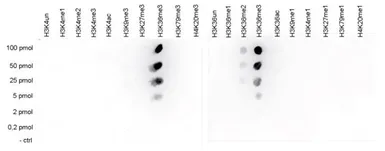
Protein Array analysis of an array containing 384 peptides with different combinations of modifications from histone H3, H4, H2A and H2B using GTX60817 Histone H3K36me3 (Tri-methy Lys36) antibody - ChIP grade. This figure shows the specificity factor, calculated as the ratio of the average intensity of all spots containing the mark, divided by the average intensity of all spots not containing the mark. The peptide array analysis shows a slight cross reaction with H4K20me3 that was not observed in dot blot.
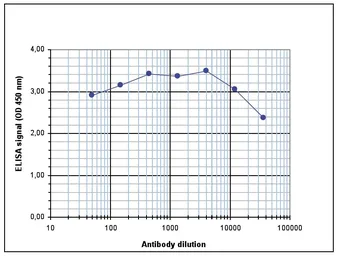
Dilution : 1:10,000
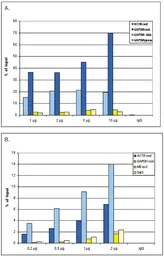
ChIP analysis of sheared chromatin from 10⁶ HeLa cells using GTX60817 Histone H3K36me3 (Tri-methy Lys36) antibody - ChIP grade. A titration consisting of 0.2, 0.5, 1 and 2 μg of antibody per ChIP experiment was analyzed. IgG (1 μg/IP) was used as a negative IP control. Quantitative PCR was performed with primers for the coding region of the active GAPDH and ACTB genes, used as positive controls, and for the coding region of the inactive MB gene and the Sat satellite repeat, used as negative controls. This figure shows the recovery, expressed as a % of input (the relative amount of immunoprecipitated DNA compared to input DNA after qPCR analysis).